Hemoglobin Cooperativity
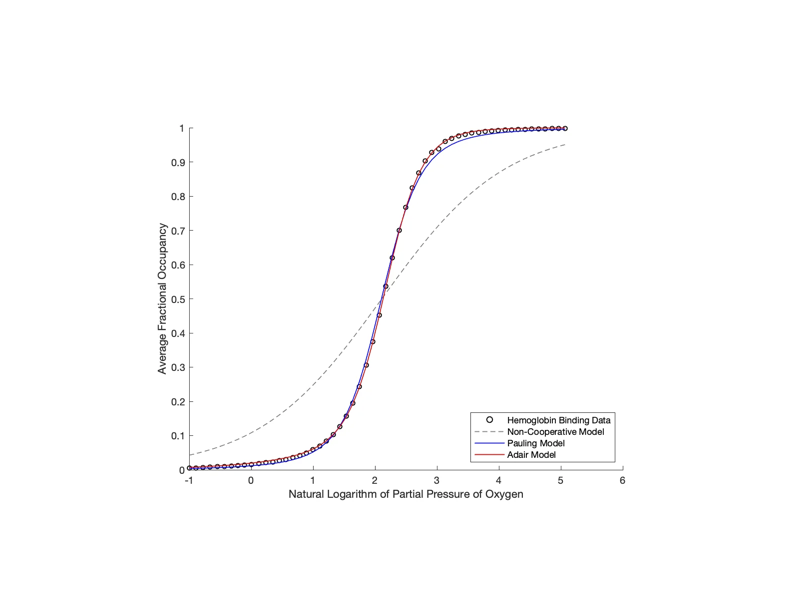
Using MATLAB, I did data analysis on experimental hemoglobin binding data. I curve-fit different models to the data, including non-cooperative, Pauling, and Adair models for hemoglobin cooperativity. Fitting the non-cooperative and Pauling models involved exploring minimal errors in a parameter space via contour plots, utilizing a brute force method, and picking the approximate minimum error value. For the Adair model, I ran a Monte Carlo simulation to fit the necessary model parameters, since there were four of them. I derived these models from scratch from thermodynamical laws and statistical mechanics, including the Grand Partition function.
Below is the poster presentation given on this project:
About
Hemoglobin proteins are found in the blood and are known for transporting oxygen. Each individual hemoglobin protein has four binding sites for oxygen. At any one time, a hemoglobin protein can have any permutation of oxygen molecules bound to it, including being empty (no oxygen molecules bound).
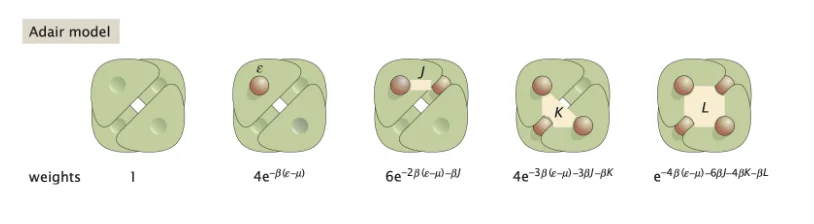
Oxygen binding sites and options in hemoglobin molecules (Phillips et. al.).
In the above image, we have regions \(J\), \(K\), and \(L\), which represent the cooperativity between oxygen molecules. From this we can assume \(J=K=L=0\) for non-cooperativity and \(K=L=0\) for the Pauling model.
Derivation
There are a few main equations that go into solving this system, which are noted in the poster. The derivation/statement of those equations are given in the following PDF:
